I'm a huge supporter of the Free and Open Source Software movement. I've written more about R than anything else on this blog, all the code I post here is free and open-source, and a while back I invited you to steal this blog under a cc-by-sa license.
Every now and then, however, something comes along that just might be worth paying for. As a director of a bioinformatics core with a very small staff, I spend a lot of time balancing costs like software licensing versus personnel/development time, so that I can continue to provide a fiscally sustainable high-quality service.
As you've likely noticed from my more recent blog/twitter posts, the core has been doing a lot of gene expression and RNA-seq work. But recently had a client who wanted to run a fairly standard case-control GWAS analysis on a dataset from dbGaP. Since this isn't the focus of my core's service, I didn't want to invest the personnel time in deploying a GWAS analysis pipeline, downloading and compiling all the tools I would normally use if I were doing this routinely, and spending hours on forums trying to remember what to do with procedural issues such as which options to specify when running EIGENSTRAT or KING, or trying to remember how to subset and LD-prune a binary PED file, or scientific issues, such as whether GWAS data should be LD-pruned at all before doing PCA.
Golden Helix
A year ago I wrote a post about the "Hitchhiker's Guide to Next-Gen Sequencing" by Gabe Rudy, a scientist at Golden Helix. After reading this and looking through other posts on their blog, I'm confident that these guys know what they're doing and it would be worth giving their product a try. Luckily, I had the opportunity to try out their SNP & Variation Suite (SVS) software (I believe you can also get a free trial on their website).
I'm not going to talk about the software - that's for a future post if the core continues to get any more GWAS analysis projects. In summary - it was fairly painless to learn a new interface, import the data, do some basic QA/QC, run a PCA-adjusted logistic regression, and produce some nice visualization. What I want to highlight here is the level of support and documentation you get with SVS.
Documentation
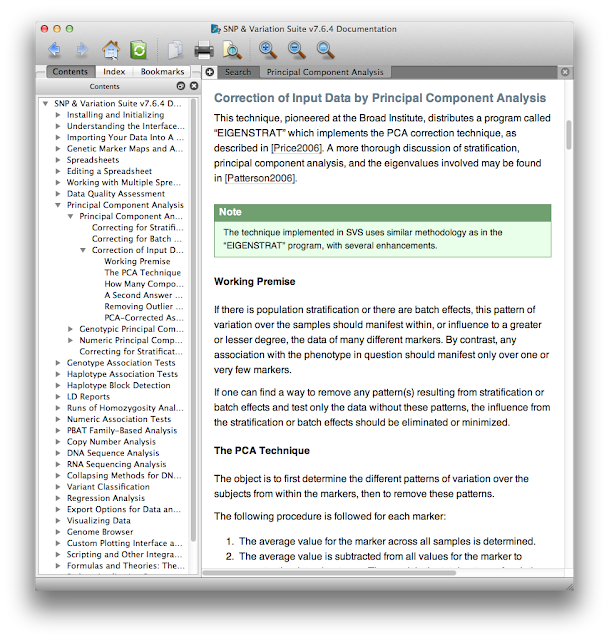
First, the documentation. At each step from data import through analysis and visualization there's a help button that opens up the manual at the page you need. This contextual manual not only gives operational details about where you click or which options to check, but also gives scientific explanations of why you might use certain procedures in certain scenarios. Here's a small excerpt of the context-specific help menu that appeared when I asked for help doing PCA.
What I really want to draw your attention to here is that even if you don't use SVS you can still view their manual online without registering, giving them your e-mail, or downloading any trialware. Think of this manual as an always up-to-date mega-review of GWAS - with it you can learn quite a bit about GWAS analysis, quality control, and statistics. For example, see this section on haplotype frequency estimation and the EM algorithm. The section on the mathematical motivation and implementation of the Eigenstrat PCA method explains the method perhaps better than the Eigenstrat paper and documentation itself. There are also lots of video tutorials that are helpful, even if you're not using SVS. This is a great resource, whether you're just trying to get a better grip on what PLINK is doing, or perhaps implementing some of these methods in your own software.
Support
Next, the support. After installing SVS on both my Mac laptop and the Linux box where I do my heavy lifting, one of the product specialists at Golden Helix called me and walked me through every step of a GWAS analysis, from QC to analysis to visualization. While analyzing the dbGaP data for my client I ran into both software-specific procedural issues as well as general scientific questions. If you've ever asked a basic question on the R-help mailing list, you know need some patience and a thick skin for all the RTFM responses you'll get. I was able to call the fine folks at Golden Helix and get both my technical and scientific questions answered in the same day. There are lots of resources for getting your questions answered, such as SEQanswers, Biostar, Cross Validated, and StackOverflow to name a few, but getting a forum response two days later from "SeqGeek96815" doesn't compare to having a team of scientists, statisticians, programmers, and product specialists on the other end of a telephone whose job it is to answer your questions.
Final Thoughts
This isn't meant to be a wholesale endorsement of Golden Helix or any other particular software company - I only wanted to share my experience stepping outside my comfortable open-source world into the walled garden of a commercially-licensed software from a for-profit company (the walls on the SVS garden aren't that high in reality - you can import and export data in any format imaginable). One of the nice things about command-line based tools is that it's relatively easy to automate a simple (or at least well-documented) process with tools like Galaxy, Taverna, or even by wrapping them with perl or bash scripts. However, the types of data my clients are collecting and the kinds of questions they're asking are always a little new and different, which means I'm rarely doing the same exact analysis twice. Because of the level of documentation and support provided to me, I was able to learn a new interface to a set of familiar procedures and run an analysis very quickly and without spending hours on forums figuring out why a particular program is seg-faulting. Will I abandon open-source tools like PLINK for SVS, Tophat-Cufflinks for CLC Workbench, BWA for NovoAlign, or R for Stata? Not in a million years. I haven't talked to Golden Helix or some of the above-mentioned companies about pricing for their products, but if I can spend a few bucks and save the time it would taken a full time technician at $50k+/year to write a new short read aligner or build a new SNP annotation database server, then I'll be able to provide a faster, high-quality, fiscally sustainable service at a much lower price for the core's clients, which is all-important in a time when federal funding is increasingly harder to come by.
Getting Genetics Done by Stephen Turner is licensed under a Creative Commons Attribution-NonCommercial 3.0 Unported License.
I caught your very thoughtful post through R-bloggers. Maybe we've crossed a threshold, when stepping outside one's comfort zone means going from open-source to commercial software, and not the other way around. (I'm finding myself there too, as after 20 years with Linux it's now Windows that confuses me.)
ReplyDeleteI don't do genetic analyses (I will learn it one of these days!) though like you I run a core analysis service. In my statistical work I find myself going back and forth between the two worlds without thinking about it. But when I had a data export question about MicroBrightField's Stereo Investigator (which my pathologist colleagues use to measure neurons) it was a pleasure to just call and get someone who knows the program in great depth and can call on the company's resources to request clarification or even new features.
I think your real step outside your comfort zone has been to point out to a sometimes religiously anti-commercial audience that, yes, paying for the polish and support can, at times, be well worth it!
-- Michael Flory
Interesting post, I think what you're doing makes a lot of sense given how much time it can take to put together the scripts required to run GWAS analysis. One thing I noticed while looking over the Golden Helix documentation that you may want to watch out for is that SVS uses PCA's to control for batch effects. Including several PCA's will probably attenuate the effects of batch, but it usually doesn't remove it very well. I include subject batch as covariate, and several of the batches are almost always highly significant at predicting phenotype. It's great that they have such a user friendly package, just make sure it's doing what you think it is.
ReplyDelete-Matthew A. Simonson
Agreed, PCA is not a cure-all, and we also use regression approaches in association tests to account for confounders. At Golden Helix we've come to the conclusion that most of the batch effect correction approaches are just band-aids and not a cure for the cause: bad experimental design. A popular blog on this topic is: Stop Ignoring Experimental Design (or my head will explode). I recently co-authored a Biostatistics editorial describing the systemic causes of the problem, and gave a webinar on the topic called Learning from our GWAS Mistakes.
DeleteA challenge with regression approaches to fixing batch effects in experiments such as GWAS is that the biggest batch effects are at the plate level and you might have 100 plates to somehow construct dummy variables for in a regression model if you flubbed your experimental design. If a block randomized design is used across plates blocking on phenotype and known sources of variability in sample collection, then you can regress out the sample collection batch effects and not have it be further exacerbated by plate effects.
Thanks for the comments. @Matthew - one of the nice things about SVS is that it tells you exactly what it's doing. I worked with the EMERGE network years ago and saw very similar things with PCs and batch variables when we tried merging similar cohorts. PCs or not, including a batch variable can't hurt.
ReplyDeleteGreat post, I love Linux and I think all the things can be done with the basic GNU tools that comes in the installation. You see in my case I started out this way then I quickly realized that I had a year to come up with a working pipeline and I didn't have staff to help with the programming so I decided to go for the licensed software for two reasons. The work I will spend is already done for me and the support. You see investigators often underestimate the amount of time required which is another reason. So time constraints with low staff and cost are two big reasons to use licensed software.
ReplyDeleteProfessor Turner what is your view on cloud computing for conducting bioinformatics analysis? Also being a core lab, do you ever see a bottle neck in the amount of samples you have to process and analyze any given day? Is turn around time for analyzed data such a an alignment to reference important? How long does it take to run a full human genome and analyze a alignment to reference? If there is a bottle neck can it be solved by moving analysis to the cloud?
ReplyDeleteThere are lots of questions in this single comment. I'd suggest the following reading:
Delete1. J. T. Dudley, A. J. Butte, In silico research in the era of cloud computing., Nature biotechnology 28, 1181–5 (2010).
2. J. T. Dudley, Y. Pouliot, R. Chen, A. A. Morgan, A. J. Butte, Translational bioinformatics in the cloud: an affordable alternative., Genome medicine 2, 51 (2010).
3. V. A. Fusaro, P. Patil, E. Gafni, D. P. Wall, P. J. Tonellato, F. Lewitter, Ed. Biomedical Cloud Computing With Amazon Web Services, PLoS Computational Biology 7, e1002147 (2011).
4. L. D. Stein, The case for cloud computing in genome informatics., Genome biology 11, 207 (2010).